Chapter 10 Cluster Analysis
10.1 Why Cluster Analysis Matters in Marketing
Cluster analysis is one of the most widely used tools in marketing analytics for market segmentation. Unlike regression or classification models, cluster analysis is an unsupervised learning technique: there is no dependent variable. Instead, the objective is to identify groups of customers who are similar to one another and meaningfully different from customers in other groups.
In marketing contexts, cluster analysis is commonly used to:
- Identify distinct customer segments
- Support targeting and positioning decisions
- Inform product design and messaging strategy
- Provide structure for downstream analysis and managerial reporting
Because cluster analysis does not optimize a predictive objective, interpretation and managerial judgment play a central role in determining whether a solution is useful.
10.2 The ffseg Dataset
This chapter uses the ffseg dataset, which contains survey responses related to fast-food consumption behaviors, attitudes, and demographics. Each row represents an individual consumer.
The dataset includes:
- Numeric attitudinal and behavioral variables suitable for segmentation
- Demographic and categorical variables used for describing clusters after they are formed
10.3 Hierarchical Agglomerative Clustering
10.3.1 Conceptual Overview
Hierarchical agglomerative clustering builds clusters from the bottom up. Each observation starts as its own cluster, and clusters are successively merged based on similarity until all observations belong to a single cluster.
Key features of hierarchical clustering include:
- No need to specify the number of clusters in advance
- A dendrogram that visualizes the clustering process
- Strong transparency for exploratory segmentation
10.3.2 Initial Hierarchical Clustering Fit
We begin by fitting a hierarchical clustering model using selected segmentation variables from the ffseg dataset. While base R can do this for us in a number of steps, we can more easily do the initial fit using the easy_hc_fit() function from the MKT4320BGSU package.
Usage:
easy_hc_fit(data, vars, dist = c("euc", "euc2", "max", "abs", "bin"),
method = c("ward", "single", "complete", "average"), k_range = 1:10,
standardize = TRUE, show_dend = TRUE, dend_max_n = 300)- where:
datais a data frame containing the full dataset.varsis a character vector of numeric segmentation variable names.distis a distance measure to use. One of: “euc”, “euc2”, “max”, “abs”, “bin”.methodis a linkage method to use. One of: “ward”, “single”, “complete”, “average”.k_rangeis an integer vector of cluster solutions to evaluate (default 1:10; allowed 1–20).standardizeis logical; if TRUE (default), standardizes segmentation variables prior to clustering.show_dendis logical; if TRUE (default), plots a dendrogram (skipped if n > dend_max_n).dend_max_nis an integer for the maximum sample size for drawing a dendrogram (default 300).
When using the easy_hc_fit() function, the results should be saved to an object. The $stop diagnostics table saved in the object provides the necessary results to help decide on a (potential) final solution. The diagnostics table summarizes multiple stopping rules and balance checks across different values of \(k\). No single statistic should be used in isolation when choosing the number of clusters. You can also view the $size_prop table to see the cluster sizes for all potential solutions.
hc_vars <- c("quality", "price", "healthy", "variety", "speed")
hc_fit <- easy_hc_fit(data = ffseg, vars = hc_vars, dist = "euc",
method = "ward", k_range = 2:8)Skipping dendrogram (n > dend_max_n).Cluster Diagnostics (Euclidean Distance / Ward's D Linkage) | ||||||
|---|---|---|---|---|---|---|
Clusters | Duda.Hart | pseudo.t2 | Silhouette | Small.Prop | Large.Prop | CV |
2 | 0.837 | 88.38 | 0.1774 | 0.3949 | 0.6051 ◊ | 0.297 * |
3 | 0.844 | 54.74 | 0.1558 | 0.2247 | 0.3949 | 0.283 * |
4 | 0.849 | 50.52 | 0.1174 | 0.0465^ | 0.3803 | 0.605 ** |
5 | 0.745 | 58.58 | 0.0945 | 0.0465^ | 0.3484 | 0.557 ** |
6 | 0.863 | 41.36 | 0.1063 | 0.0465^ | 0.3484 | 0.640 ** |
7 | 0.822 | 36.11 | 0.1065 | 0.0465^ | 0.2460 | 0.497 * |
8 | 0.821 | 40.00 | 0.1156 | 0.0465^ | 0.2460 | 0.468 * |
^ Smallest cluster < 5% of sample. | ||||||
◊ Largest cluster > 50% of sample. | ||||||
* CV < 0.5 well balanced; ** moderately imbalanced. | ||||||
Cluster Size Proportions by Solution | ||||||||
|---|---|---|---|---|---|---|---|---|
Solution | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | Cluster 5 | Cluster 6 | Cluster 7 | Cluster 8 |
2-cluster solution | 0.3949 | 0.6051 | ||||||
3-cluster solution | 0.3949 | 0.3803 | 0.2247 | |||||
4-cluster solution | 0.3484 | 0.3803 | 0.2247 | 0.0465 | ||||
5-cluster solution | 0.3484 | 0.2301 | 0.2247 | 0.1503 | 0.0465 | |||
6-cluster solution | 0.3484 | 0.1290 | 0.1011 | 0.2247 | 0.1503 | 0.0465 | ||
7-cluster solution | 0.2460 | 0.1290 | 0.1024 | 0.1011 | 0.2247 | 0.1503 | 0.0465 | |
8-cluster solution | 0.2460 | 0.1290 | 0.1024 | 0.1011 | 0.1356 | 0.1503 | 0.0891 | 0.0465 |
10.3.3 Selecting the Number of Clusters
When selecting the number of clusters, consider:
- Statistical indicators such as the Gap statistic
- Whether clusters are reasonably balanced in size
- Whether the resulting segments are interpretable and actionable
Extremely small clusters or one dominant cluster are often warning signs in marketing segmentation.
10.3.4 Final Hierarchical Clustering Solution
Once a reasonable number of clusters has been selected, we finalize the hierarchical clustering solution and attach cluster membership back to the full dataset using the easy_hc_final() function from the MKT4320BGSU package.
Usage:
easy_hc_final(fit, data, k, cluster_col = "cluster", conf_level = 0.95,
auto_print = TRUE)- where:
fitis the object returned byeasy_hc_fit.datais the original full dataset used to create fit.kis an integer representing the number of clusters to extract.cluster_colis the name of the cluster column to append todata(default “cluster”). Must not already exist in data.conf_levelconfidence level for CI error bars when plotting (default 0.95).auto_printis logical; if TRUE (default), prints props and profile and displays the plot (if available).
Using the example from above with a 3-cluster solution, the cluster profile table reports mean values of the segmentation variables for each cluster. These means should be interpreted in relative terms across clusters. The cluster plot visualizes those means.
hc_final <- easy_hc_final(fit = hc_fit, data = ffseg, k = 3,
cluster_col = "hc_cluster", auto_print = FALSE)
hc_final$props Cluster N Prop
1 1 297 0.3949468
2 2 286 0.3803191
3 3 169 0.2247340 Cluster quality price healthy variety speed
1 1 4.737374 4.050505 3.996633 3.905724 3.936027
2 2 3.723776 4.520979 2.339161 2.979021 3.674825
3 3 3.905325 3.047337 2.568047 2.952663 3.112426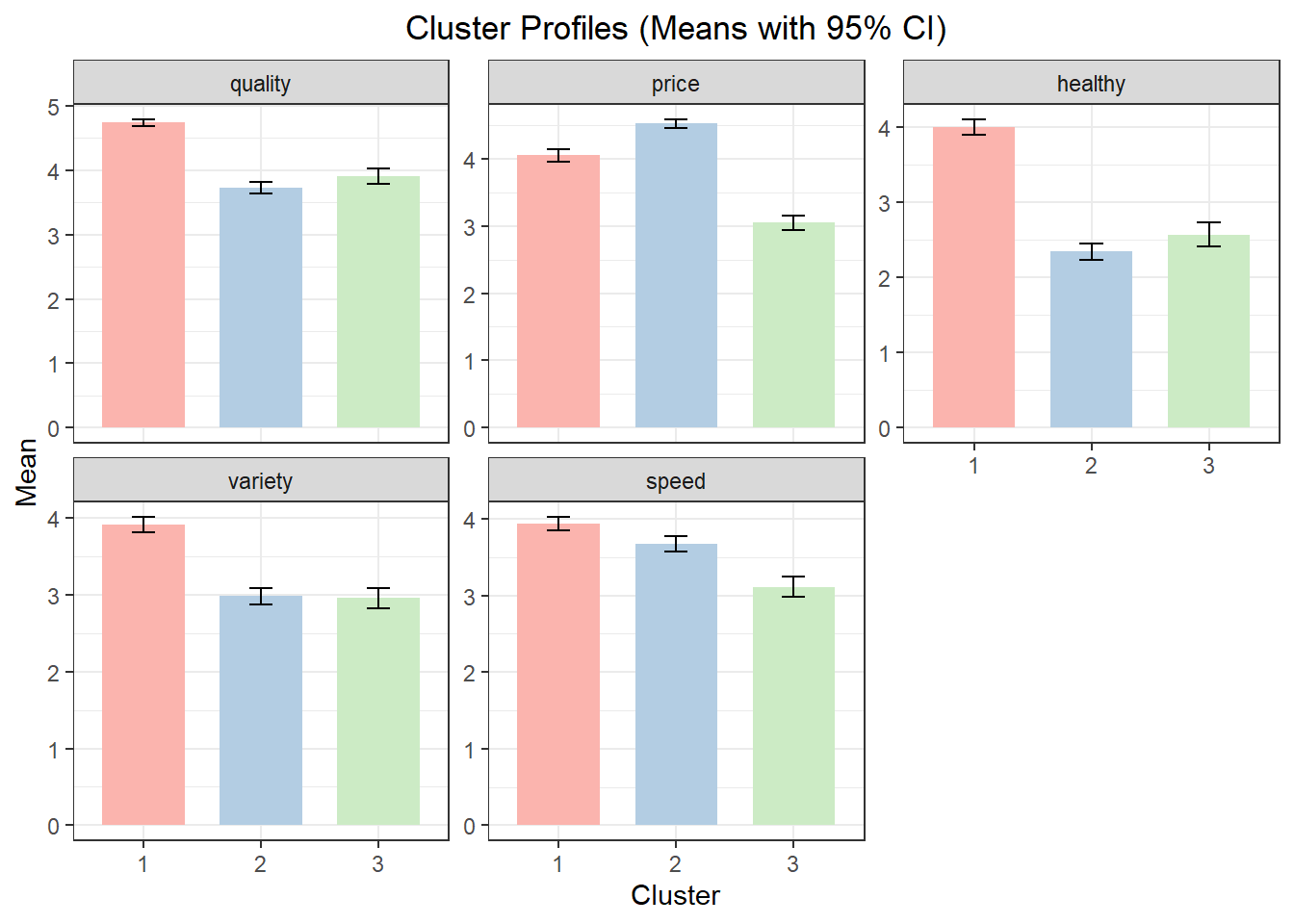
10.4 Describing and Labeling Hierarchical Clusters
Segmentation variables alone rarely provide enough context to understand who the clusters represent. Additional variables are used to describe and label the clusters. For both hierarchical clustering and k-means clustering (see below), use the easy_cluster_decribe() function from the MKT4320BGSU package to automate this process.
Usage:
easy_cluster_describe(data, cluster_col = "cluster", var, alpha = 0.05,
drop_missing = TRUE, auto_print = TRUE), digits = 4- where:
datais the data frame containing the cluster membership column and variables to describe that was saved to theeasy_hc_finalobject; should beobjectname$data.cluster_colis a character string naming the cluster membership column in data (default = “cluster”).varis a character string naming the single variable to describe.alphais the significance level for hypothesis tests (default = 0.05).drop_missingis logical; if TRUE (default), rows with missing cluster membership are dropped prior to analysis.auto_printis logical; if TRUE (default), prints summaries to the console; if FALSE returns results listdigitsis an integer for rounding/formatting of numeric output (default = 4).
Using the results from above (i.e., the hc_final$data object) , the output below highlights which variables significantly differentiate clusters and provides detailed summaries only where differences are statistically meaningful.
Variable: usertype (categorical)
Overall test
Variable Test p_value Significant
-------- ---------- ------- -----------
usertype Chi-square 0.05500 FALSE
Not significant at alpha = 0.05; no post-hoc shown.
Variable: spend (categorical)
Overall test
Variable Test p_value Significant
-------- ---------- -------- -----------
spend Chi-square 0.003500 TRUE
Cross-tab (column %; clusters are columns)
Level 1 2 3
------------ ----- ----- -----
$10 or more 25.25 17.48 26.63
$5 to $9 61.95 63.64 65.68
Less than $5 12.79 18.88 7.692
Post-hoc (Holm-adjusted p-values)
Row 1:2 1:3 2:3
------------------------ ------- ------ --------
$10 or more:$5 to $9 0.3758 1.000 0.5700
$10 or more:Less than $5 0.06200 0.5700 0.002300
$5 to $9:Less than $5 0.5700 0.5700 0.03350
Variable: hours (continuous)
Overall test
Variable Test p_value Significant
-------- ----- ------- -----------
hours ANOVA 0 TRUE
Cluster means (N, Mean, SD)
Cluster N Mean SD
------- ----- ----- ------
1 297.0 3.710 0.9428
2 286.0 3.542 1.014
3 169.0 3.065 1.070
Significant differences: 1 > 3, 2 > 3
Post-hoc comparisons (Games-Howell)
.y. group1 group2 estimate conf.low conf.high p.adj p.adj.signif
----- ------ ------ -------- -------- --------- --------------- ------------
hours 1 2 -0.1685 -0.3592 0.02224 0.09600 ns
hours 1 3 -0.6453 -0.8781 -0.4126 0.0000000007900 ****
hours 2 3 -0.4769 -0.7166 -0.2372 0.00001220 ****
Variable: eatin (continuous)
Overall test
Variable Test p_value Significant
-------- ----- ------- -----------
eatin ANOVA 0.02040 TRUE
Cluster means (N, Mean, SD)
Cluster N Mean SD
------- ----- ----- -----
1 297.0 3.986 2.000
2 286.0 4.441 1.978
3 169.0 4.160 1.903
Significant differences: 2 > 1
Post-hoc comparisons (Games-Howell)
.y. group1 group2 estimate conf.low conf.high p.adj p.adj.signif
----- ------ ------ -------- -------- --------- ------- ------------
eatin 1 2 0.4540 0.06692 0.8411 0.01700 *
eatin 1 3 0.1732 -0.2664 0.6129 0.6230 ns
eatin 2 3 -0.2808 -0.7218 0.1602 0.2930 ns
Variable: mealplan (categorical)
Overall test
Variable Test p_value Significant
-------- ---------- ------- -----------
mealplan Chi-square 0.8882 FALSE
Not significant at alpha = 0.05; no post-hoc shown.
Variable: gender (categorical)
Overall test
Variable Test p_value Significant
-------- ---------- ------- -----------
gender Chi-square 0 TRUE
Cross-tab (column %; clusters are columns)
Level 1 2 3
------ ----- ----- -----
Female 81.82 67.83 59.17
Male 18.18 32.17 40.83
Post-hoc (Holm-adjusted p-values)
Row 1:2 1:3 2:3
----------- --------- --- -------
Female:Male 0.0002000 0 0.0681010.5 K-Means Clustering
10.5.1 Conceptual Overview
k-means clustering is a partition-based method that assigns observations to a fixed number of clusters by minimizing within-cluster variation. Unlike hierarchical clustering, k-means requires the analyst to specify the number of clusters in advance.
k-means is often preferred when:
- Working with larger datasets
- Refining a solution suggested by hierarchical clustering
- Stability and computational efficiency are priorities
The process used for k-means clustering follows the process used above in hierarchical agglomerative clustering. That is, we first initially fit a number of potential solutions, and then we anlayze a final solution.
10.5.2 Initial K-Means Clustering Fit
We first fit k-means clustering solutions across a range of cluster counts using the easy_km_fit() function from the MKT4320BGSU package.
Usage:
easy_km_fit(data, vars, k_range = 1:10, standardize = TRUE, nstart = 25,
iter.max = 100, B = 20, seed = 4320)- where:
datais a data frame containing the full dataset.varsis a character vector of numeric variable names used for clustering.k_rangeis an integer vector of cluster counts to evaluate (default = 1:10; allowed values 1–20).standardizeis logical; if TRUE (default), clustering variables are standardized before fitting k-means.nstartis an integer; number of random starts for each k-means solution (default = 25).iter.maxis an integer; maximum number of iterations allowed for each k-means run (default = 100).Bis an integer; number of Monte Carlo bootstrap samples used to compute the Gap statistic (default = 20).seedis an integer; random seed for reproducible results (default = 4320).
We now fit the same variables we used above in hierarchical clustering. As before, the results should be saved to an object. The $diag diagnostics table saved in the object provides the necessary results to help decide on a (potential) final solution. The diagnostics table summarizes multiple stopping rules and balance checks across different values of \(k\). No single statistic should be used in isolation when choosing the number of clusters. You can also view the $size_prop table to see the cluster sizes for all potential solutions.
km_vars <- c("quality", "price", "healthy", "variety", "speed")
km_fit <- easy_km_fit(data = ffseg, vars = km_vars, k_range = 2:8)
km_fit$diagK-Means Cluster Diagnostics (Standardized) | |||||
|---|---|---|---|---|---|
Clusters | Silhouette | Gap.Stat | Small.Prop | Large.Prop | CV |
2 | 0.2120 | 0.6307 | 0.4761 | 0.5239 ◊ | 0.068 * |
3 | 0.1973 | 0.6150 | 0.2739 | 0.4096 | 0.208 * |
4 | 0.1610 | 0.5953 | 0.2168 | 0.2806 | 0.105 * |
5 | 0.1679 | 0.5855 | 0.1569 | 0.2340 | 0.178 * |
6 | 0.1719 | 0.5827 | 0.1303 | 0.2354 | 0.224 * |
7 | 0.1803 | 0.5741 | 0.1184 | 0.1835 | 0.195 * |
8 | 0.1804 | 0.5747 | 0.0851 | 0.1582 | 0.185 * |
^ Smallest cluster < 5% of sample. | |||||
◊ Largest cluster > 50% of sample. | |||||
* CV < 0.5 well balanced; ** moderately imbalanced. | |||||
Cluster Size Proportions by Solution | ||||||||
|---|---|---|---|---|---|---|---|---|
Solution | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | Cluster 5 | Cluster 6 | Cluster 7 | Cluster 8 |
2-cluster solution | 0.4761 | 0.5239 | ||||||
3-cluster solution | 0.3165 | 0.4096 | 0.2739 | |||||
4-cluster solution | 0.2168 | 0.2487 | 0.2540 | 0.2806 | ||||
5-cluster solution | 0.2128 | 0.1569 | 0.2340 | 0.2287 | 0.1676 | |||
6-cluster solution | 0.1423 | 0.1503 | 0.1303 | 0.1715 | 0.1702 | 0.2354 | ||
7-cluster solution | 0.1263 | 0.1184 | 0.1516 | 0.1835 | 0.1769 | 0.1210 | 0.1223 | |
8-cluster solution | 0.1330 | 0.1436 | 0.1396 | 0.1582 | 0.1117 | 0.0851 | 0.1157 | 0.1130 |
10.5.3 Final K-Means Clustering Solution
After selecting a value for \(k\), we finalize the k-means solution using the easy_km_final() function from the MKT4320BGSU package.
Usage:
easy_km_final(fit, data, k, cluster_col = "cluster", conf_level = 0.95,auto_print = TRUE)- where:
fitis an object returned byeasy_km_fit().datais the original full dataset used ineasy_km_fit().kis an integer; number of clusters to extract.cluster_colis a character string; name of the cluster column to append to data (default = “cluster”).conf_levelis numeric; confidence level for CI error bars (default = 0.95).auto_printis logical; if TRUE (default), prints selected outputs and displays the plot when the function is run.
Using the example from above with a 3-cluster solution, the cluster profile table reports mean values of the segmentation variables for each cluster. These means should be interpreted in relative terms across clusters. The cluster plot visualizes those means.
km_final <- easy_km_final(fit = km_fit, data = ffseg, k = 3,
cluster_col = "km_cluster", auto_print = FALSE)
km_final$props Cluster N Prop
1 1 238 0.3164894
2 2 308 0.4095745
3 3 206 0.2739362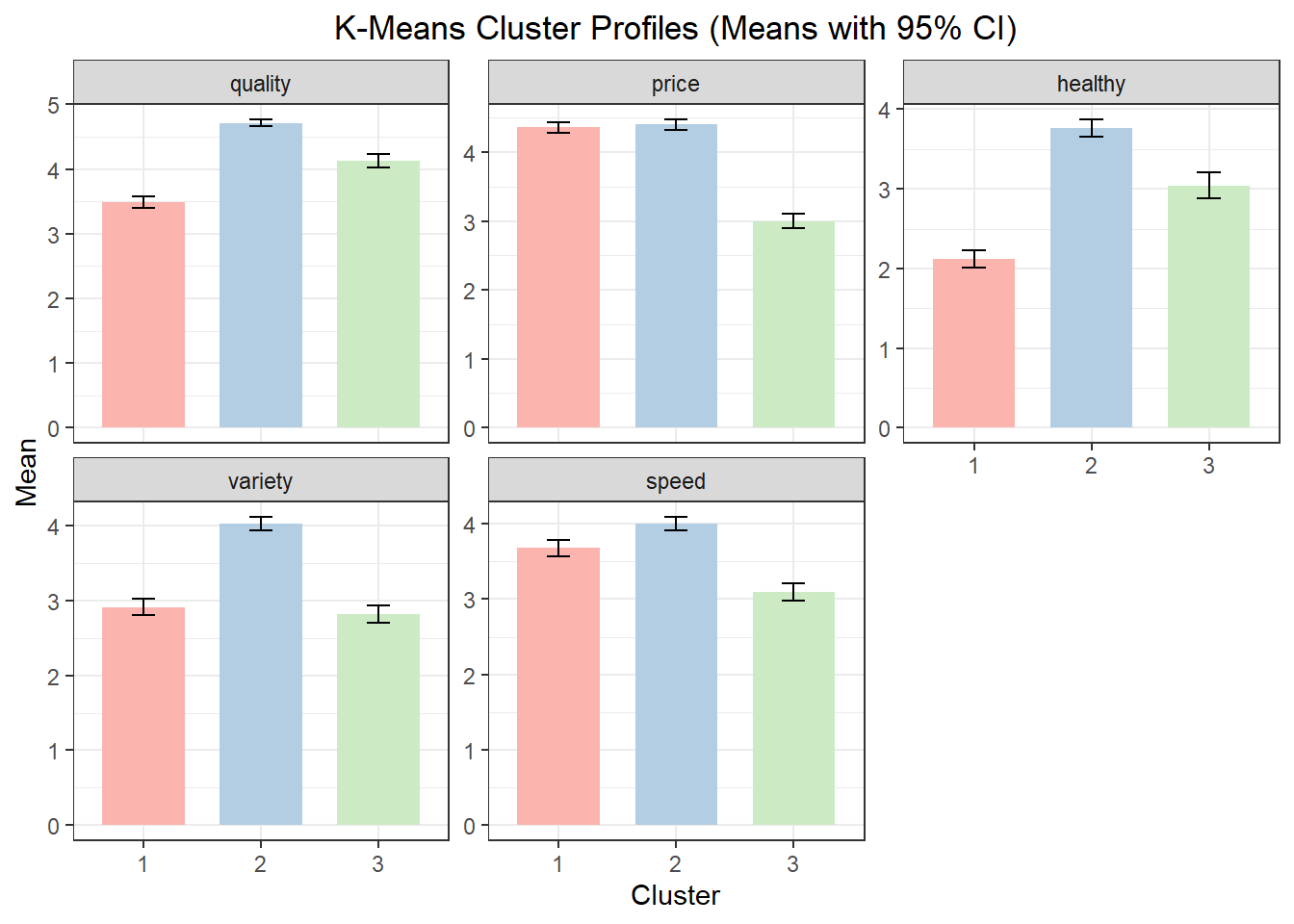
Cluster quality price healthy variety speed
1 1 3.483193 4.361345 2.117647 2.907563 3.680672
2 2 4.714286 4.399351 3.762987 4.022727 4.000000
3 3 4.131068 3.000000 3.043689 2.815534 3.09708710.6 Describing and Labeling k-Means Clusters
As with hierarchical clustering, segmentation variables alone rarely provide enough context to understand who the clusters represent. Additional variables are used to describe and label the clusters. As with hierarchical clustering, use the easy_cluster_decribe() function from the MKT4320BGSU package to automate this process. See Section 10.4
10.7 Comparing Clustering Approaches
Hierarchical and k-means clustering often produce similar but not identical segmentation results. Differences between solutions can be informative and may reveal alternative ways to think about the market.
In practice, analysts often:
- Use hierarchical clustering to explore the structure of the data
- Use k-means clustering to refine and stabilize the final solution
10.8 Chapter Summary
In this chapter, we:
- Introduced cluster analysis as a tool for marketing segmentation
- Applied hierarchical and k-means clustering to the
ffsegdataset - Used diagnostics to guide the choice of the number of clusters
- Described and interpreted clusters in a marketing context
10.9 What’s Next
In the next chapter, we shift from grouping customers to summarizing and visualizing variables. Specifically, we will introduce Principal Components Analysis (PCA) and then extend it to PCA perceptual maps. PCA is a dimensionality-reduction technique that helps simplify complex datasets by transforming many correlated variables into a smaller set of components that capture the most important patterns in the data.
You will learn how PCA can be used to:
- Reduce a large set of variables into a smaller number of interpretable dimensions
- Identify underlying structures in consumer perceptions and evaluations
- Prepare data for visualization and communication
We will then use these components to create perceptual maps, which are widely used in marketing to:
- Visualize brand or product positions
- Understand competitive structure
- Support positioning and differentiation decisions